In this post, I will show how to scrape google scholar. Particularly, we will use the 'rvest' R package to scrape the google scholar account of my PhD advisor. We will see his coauthors, how many times they have been cited and their affiliations. “rvest, inspired by libraries like beautiful soup, makes it easy to scrape (or harvest) data from html web pages”, wrote Hadley Wickham on RStudio Blog. Since it is designed to work with magrittr, we can express complex operations as elegant pipelines composed of simple and easily understood pieces of code.
Load required libraries:
We will use ggplot2 to create plots.
library(rvest) library(ggplot2)
How many times have his papers been cited
Let’s use SelectorGadget to find out which css selector matches the “cited by” column.
page <- read_html("https://scholar.google.com/citations?user=sTR9SIQAAAAJ&hl=en&oi=ao")
Specify the css selector in html_nodes() and extract the text with html_text(). Finally, change the string to numeric using as.numeric().
citations <- page %>% html_nodes ("#gsc_a_b .gsc_a_c") %>% html_text()%>%as.numeric()
See the number of citations:
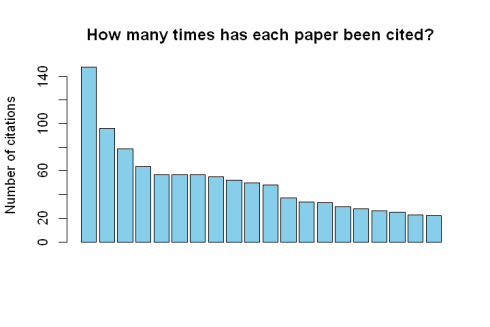
citations 148 96 79 64 57 57 57 55 52 50 48 37 34 33 30 28 26 25 23 22
Create a barplot of the number of citation:
barplot(citations, main="How many times has each paper been cited?", ylab='Number of citations', col="skyblue", xlab="")
Coauthors, thier affilations and how many times they have been cited
My PhD advisor, Ben Zaitchik, is a really smart scientist. He not only has the skills to create network and cooperate with other scientists, but also intelligence and patience.
Next, let’s see his coauthors, their affiliations and how many times they have been cited.
Similarly, we will use SelectorGadget to find out which css selector matches the Co-authors.
page <- read_html("https://scholar.google.com/citations?view_op=list_colleagues&hl=en&user=sTR9SIQAAAAJ")
Coauthors = page%>% html_nodes(css=".gsc_1usr_name a") %>% html_text()
Coauthors = as.data.frame(Coauthors)
names(Coauthors)='Coauthors'
Now let’s exploring Coauthors
head(Coauthors)
dim(Coauthors)
Coauthors
1 Jason Evans
2 Mutlu Ozdogan
3 Rasmus Houborg
4 M. Tugrul Yilmaz
5 Joseph A. Santanello, Jr.
6 Seth Guikema
[1] 27 1
As of today, he has published with 27 people.
How many times have his coauthors been cited?
page <- read_html("https://scholar.google.com/citations?view_op=list_colleagues&hl=en&user=sTR9SIQAAAAJ")
citations = page%>% html_nodes(css = ".gsc_1usr_cby")%>%html_text()
citations
[1] "Cited by 2231" "Cited by 1273" "Cited by 816" "Cited by 395" "Cited by 652" "Cited by 1531"
[7] "Cited by 674" "Cited by 467" "Cited by 7967" "Cited by 3968" "Cited by 2603" "Cited by 3468"
[13] "Cited by 3175" "Cited by 121" "Cited by 32" "Cited by 469" "Cited by 50" "Cited by 11"
[19] "Cited by 1187" "Cited by 1450" "Cited by 12407" "Cited by 1939" "Cited by 9" "Cited by 706"
[25] "Cited by 336" "Cited by 186" "Cited by 192"
Let’s extract the numeric characters only using global substitute.
citations = gsub('Cited by','', citations)
citations
[1] " 2231" " 1273" " 816" " 395" " 652" " 1531" " 674" " 467" " 7967" " 3968" " 2603" " 3468" " 3175"
[14] " 121" " 32" " 469" " 50" " 11" " 1187" " 1450" " 12407" " 1939" " 9" " 706" " 336" " 186"
[27] " 192"
Change string to numeric and then to data frame to make it easy to use with ggplot2
citations = as.numeric(citations) citations = as.data.frame(citations)
Affilation of coauthors
page <- read_html("https://scholar.google.com/citations?view_op=list_colleagues&hl=en&user=sTR9SIQAAAAJ")
affilation = page %>% html_nodes(css = ".gsc_1usr_aff")%>%html_text()
affilation = as.data.frame(affilation)
names(affilation)='Affilation'
Now, let’s create a data frame that consists of coauthors, citations and affilations
cauthors=cbind(Coauthors, citations, affilation)
cauthors
Coauthors citations Affilation
1 Jason Evans 2231 University of New South Wales
2 Mutlu Ozdogan 1273 Assistant Professor of Environmental Science and Forest Ecology, University of Wisconsin
3 Rasmus Houborg 816 Research Scientist at King Abdullah University of Science and Technology
4 M. Tugrul Yilmaz 395 Assistant Professor, Civil Engineering Department, Middle East Technical University, Turkey
5 Joseph A. Santanello, Jr. 652 NASA-GSFC Hydrological Sciences Laboratory
.....
Re-order coauthors based on their citations
Let’s re-order coauthors based on their citations so as to make our plot in a decreasing order.
cauthors$Coauthors <- factor(cauthors$Coauthors, levels = cauthors$Coauthors[order(cauthors$citations, decreasing=F)])
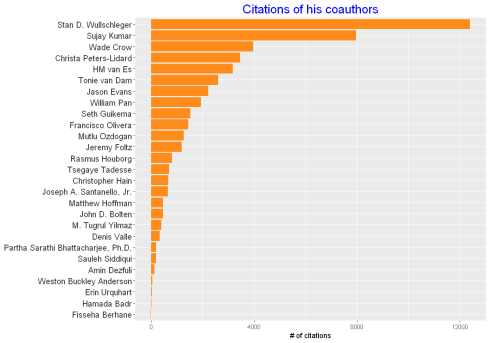
ggplot(cauthors,aes(Coauthors,citations))+geom_bar(stat="identity", fill="#ff8c1a",size=5)+
theme(axis.title.y = element_blank())+ylab("# of citations")+
theme(plot.title=element_text(size = 18,colour="blue"), axis.text.y = element_text(colour="grey20",size=12))+
ggtitle('Citations of his coauthors')+coord_flip()
He has published with scientists who have been cited more than 12000 times and with students like me who are just toddling.
Summary
In this post, we saw how to scrape Google Scholar. We scraped the account of my advisor and got data on the citations of his papers and his coauthors with thier affilations and how many times they have been cited.
As we have seen in this post, it is easy to scrape an html page using the rvest R package. It is also important to note that SelectorGadget is useful to find out which css selector matches the data of our interest.
Update: My advisor told me that Google Scholar picks up a minority of his co-authors. Some of the scientists who published with him and who my advisor would expect to be the most cited don’t show up. Further, the results for some others are counterintuitive (e.g., seniors who have more publications, have less Google Scholar citations than their juniors). So, Google Scholar data should be used with caution.
If you have any question feel free to post a comment below.